How to sort chromosomes in Golden Helix Genome Browser?
asked 2017-10-15 18:35:10 -0600
This post is a wiki. Anyone with karma >75 is welcome to improve it.
Dear,
I came across a problem that I don't know how to solve. I downloaded the new version of rainbow trout genome from NCBI website. I manually uploaded the FNA file of the genome into the Genome Browser. After I created the genome in the Genome Browse, the order of chromosomes was messed up. What shall I do to fix it?
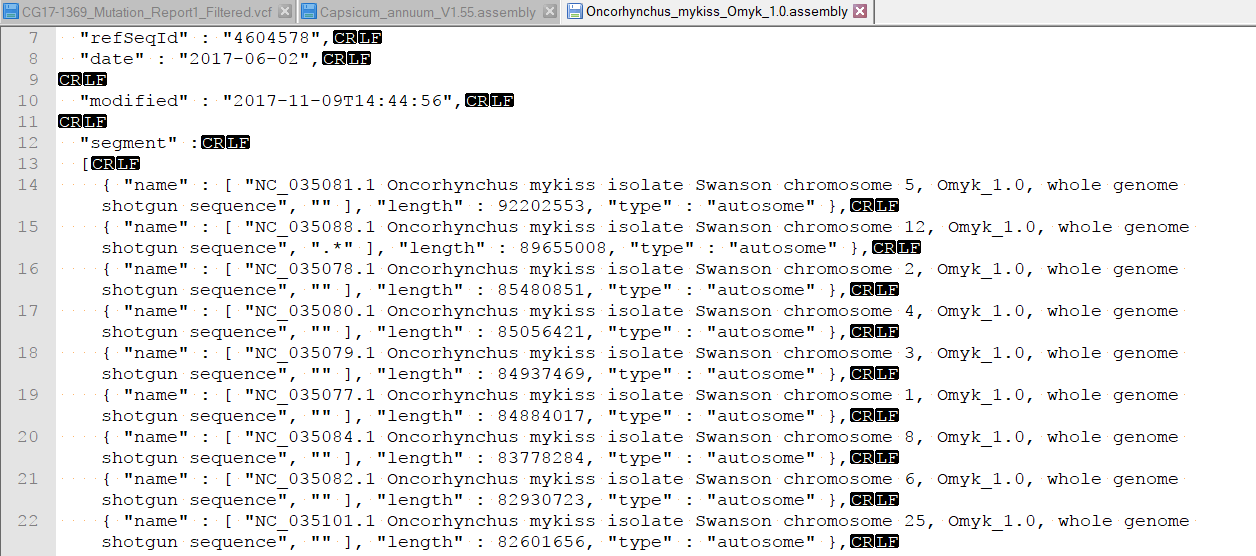
My second question is that I uploaded a vcf file to the Genome Browser. Due to the different order of chromosomes between the created genome and vcf file, the Genome Browser cannot match up. Attached is the screenshot of the problem. I would be highly appreciated if you could help to solve the problems. Thank you very much.
Shawn